IMP cyclohydrolase
| IMP cyclohydrolase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC number | 3.5.4.10 | ||||||||
| CAS number | 9013-81-4 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / EGO | ||||||||
| |||||||||
| IMP cyclohydrolase-like protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
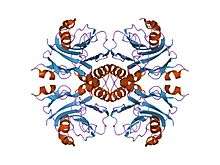 crystal structure of puro from methanothermobacter thermoautotrophicus | |||||||||
| Identifiers | |||||||||
| Symbol | IMP_cyclohyd | ||||||||
| Pfam | PF07826 | ||||||||
| InterPro | IPR020600 | ||||||||
| |||||||||
In enzymology, an IMP cyclohydrolase (EC 3.5.4.10) is an enzyme that catalyzes the chemical reaction
- IMP + H2O 5-formamido-1-(5-phospho-D-ribosyl)imidazole-4-carboxamide
Thus, the two substrates of this enzyme are IMP and H2O, whereas its product is 5-formamido-1-(5-phospho-D-ribosyl)imidazole-4-carboxamide.
This enzyme belongs to the family of hydrolases, those acting on carbon-nitrogen bonds other than peptide bonds, specifically in cyclic amidines. The systematic name of this enzyme class is IMP 1,2-hydrolase (decyclizing). Other names in common use include inosinicase, and inosinate cyclohydrolase. This enzyme catalyses the cyclisation of 5-formylamidoimidazole-4-carboxamide ribonucleotide to IMP, a reaction which is important in de novo purine biosynthesis in archaeal species.[1]
Structural studies
In most cases this single-domain protein is arranged to form an overall fold that consists of a four-layered alpha-beta-beta-alpha core structure. The two antiparallel beta-sheets pack against each other and are covered by alpha-helices on one face of the molecule. The protein is structurally similar to members of the N-terminal nucleophile (NTN) hydrolase superfamily. A deep pocket was in fact found on the surface of IMP cyclohydrolase in a position equivalent to that of active sites of NTN-hydrolases, but an N-terminal nucleophile could not be found. Therefore, it is thought that this enzyme is structurally but not functionally similar to members of the NTN-hydrolase family.[2]
As of late 2007, 14 structures have been solved for this class of enzymes, with PDB accession codes 1G8M, 1M9N, 1OZ0, 1P4R, 1PKX, 1PL0, 1THZ, 2B1G, 2B1I, 2IU0, 2IU3, 2NTK, 2NTL, and 2NTM.
References
- ↑ Graupner M, Xu H, White RH (March 2002). "New class of IMP cyclohydrolases in Methanococcus jannaschii". J. Bacteriol. 184 (5): 1471–3. doi:10.1128/jb.184.5.1471-1473.2002. PMC 134845
 . PMID 11844782.
. PMID 11844782. - ↑ Saridakis V, Christendat D, Thygesen A, Arrowsmith CH, Edwards AM, Pai EF (July 2002). "Crystal structure of Methanobacterium thermoautotrophicum conserved protein MTH1020 reveals an NTN-hydrolase fold". Proteins. 48 (1): 141–3. doi:10.1002/prot.10147. PMID 12012346.
Further reading
- FLAKS JG, ERWIN MJ, BUCHANAN JM (1957). "Biosynthesis of the purines. XVIII 5-Amino-1-ribosyl-4-imidazolecarboxamide 5'-phosphate transformylase and inosinicase". J. Biol. Chem. 229 (2): 603–12. PMID 13502325.
This article incorporates text from the public domain Pfam and InterPro IPR020600