DHRS7B
| DHRS7B | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 |
|||||||||||||||||
| Identifiers | |||||||||||||||||
| Aliases | DHRS7B, SDR32C1, CGI-93, dehydrogenase/reductase (SDR family) member 7B, dehydrogenase/reductase 7B | ||||||||||||||||
| External IDs | MGI: 2384931 HomoloGene: 41044 GeneCards: DHRS7B | ||||||||||||||||
| |||||||||||||||||
| Orthologs | |||||||||||||||||
| Species | Human | Mouse | |||||||||||||||
| Entrez | |||||||||||||||||
| Ensembl | |||||||||||||||||
| UniProt | |||||||||||||||||
| RefSeq (mRNA) | |||||||||||||||||
| RefSeq (protein) | |||||||||||||||||
| Location (UCSC) | Chr 17: 21.12 – 21.19 Mb | Chr 11: 60.83 – 60.86 Mb | |||||||||||||||
| PubMed search | [1] | [2] | |||||||||||||||
| Wikidata | |||||||||||||||||
| View/Edit Human | View/Edit Mouse |
Dehydrogenase/reductase (SDR family) member 7B is an enzyme encoded by the DHRS7B gene in humans, found on chromosome 17p11.2.[3] DHRS7B encodes a protein that is predicted to function in steroid hormone regulation.[4][5][6] A deletion in the chromosomal region 17p11.2 has been associated with Smith-Magenis Syndrome, a genetic developmental disorder.[7]
Gene
Overview
The DHRS7B gene is located on the positive strand of chromosome 17, beginning at position 21030258 and ending at position 21094836 (64579 bp).[8] DHRS7B contains seven exons with no predicted alternate splice forms, resulting in an 1841 bp mRNA product.[8][9]
Upstream of DHRS7B on the negative strand of chromosome 17p11.2 are the genes Coiled-coil domain containing 144 family, N-terminal-like (CCDC144NL) and Ubiquitin specific peptidase 22 (USP22).[10] Downstream of DHSRS7B on the negative strand of chromosome 17p11.2 is the gene Transmembrane protein 11 (TMEM11), and on the positive strand is the gene Mitogen-activated protein kinase, kinase 3 (MAP2K3).[10]
Gene expression
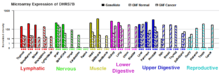
Microarray and EST data indicates that the DHRS7B gene is highly expressed in the testes, thyroid, kidneys, and adipose tissues. There is moderate expression in the brain, pancreas, mammary glands, and ovaries. Finally, there is little expression in spleen, thymus, tonsils, bone marrow, and bladder.[11][12]
Protein structure
The DHRS7B gene has a predicted protein product that is 325 amino acids, a molecular weight of 35.1 kDa, and an isoelectric point of 9.867.[13][14] There is one predicted transmembrane domain in the protein sequence, a large neutrally charged region spanning residues 18-38.[13][14] No signal peptides have been identified in DHRS7B; cellular localization remains unclear.[15]
DHRS7B is a member of the short chain dehydrogenase/reductase (SDR) superfamily and possesses characteristic features of an SDR within the protein sequence. The following table identifies sequences in the protein and corresponding function.[16]
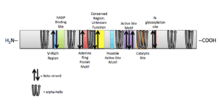
| Sequence | Function |
|---|---|
| "VVV" | Valine-rich region, unknown function |
| "TGXXXGXG" | NADP binding site |
| "NXXG" | Possible active site motif |
| "DXXD" | Adenine ring pocket motif |
| "GXXXXXSS" | Possible active site motif |
| "SXYXXXK" | Catalytic site, with upstream serine residue |
| "LXNNXG" | Conserved region, unknown function |
| "NLS" | N-glycosylation site |
Interactions
In humans, DHRS7B has been shown to physically interact with other proteins such as Mediator complex subunit 19 (MED19) and Brain and reproductive expressed-modulator protein (BRE).[17] MED19 was found to interact with DHRS7B through a two hybrid screening approach and plays a role as a co-activator in regulated transcription of most RNA polymerase II dependent genes.[18] BRE is a component of the BRCA1-A complex, which recognizes Lys-63 linked ubiquitinated histones H2A and H2AX DNA lesion sites (identified using anti-tag coimmunoprecipitation).[19] Other proteins interacting with DHRS7B have only been identified through text-mining.
Homology
Orthologs
Conservation of the DHRS7B protein sequence has been observed highly in mammals; moderately in reptiles, birds, fish and amphibians; minimally in invertebrates, insects, and fungi.[20]
Paralogs
Paralogs of DHRS7B are all in the SDR superfamily and conservation of the SDR functional motifs was identified in a multiple sequence alignment.[20][21]
| Common Name | Accession # | Sequence Length | Sequence Identity | Sequence Similarity |
|---|---|---|---|---|
| DHRS7B | NP_056325.2 | 325 aa | 100% | 100% |
| DHRS7C | AA_147025.1 | 308 aa | 46% | 66% |
| DHRS7 | CAH56402 | 375 aa | 37% | 53% |
| HBD1 | AAA58352 | 343 aa | 35% | 56% |
| RDH8 | EAW84062 | 331 aa | 34% | 53% |
| RDH16 | AAC39922 | 317 aa | 33% | 53% |
| KDSR | NP002026 | 332 aa | 31% | 51% |
| HSD11B1 | AAK83653 | 292 aa | 29% | 50% |
| DHRS9 | AAH58883 | 319 aa | 29% | 50% |
| RDH5 | AAH28298 | 318 aa | 30% | 49% |
Clinical significance
DHRS7B has been identified in the Smith-Magenis Syndrome region, where a deletion in this chromosomal region (17p11.2) causes a genetic developmental disorder.[6] In breast cancer cells expressing CD44 and CD24, DHRS7B expression was observed to be down regulated.[22] CD44 is an antigen found on the surface of most cell types and functions as a receptor that binds tissue macromolecules. Additionally, it acts as an adhesion molecule for leukocytes on peripheral lymphoid organs and inflammation sites. CD24 is associated with B-cells, epithelial cells, and dendritic cells, functioning as an adhesion molecule and shown to enhance a tumor cell's capability of metastasizing.[23]
References
- ↑ "Human PubMed Reference:".
- ↑ "Mouse PubMed Reference:".
- ↑ "Entrez Gene: Dehydrogenase/reductase (SDR family) member 7B".
- ↑ "Genecards: DHRS7B Gene protein-coding GIFtS 47".
- ↑ Tannin GM, Agarwal AK, Monder C, New MI, White PC (September 1991). "The human gene for 11 beta-hydroxysteroid dehydrogenase. Structure, tissue distribution, and chromosomal localization". J. Biol. Chem. 266 (25): 16653–8. PMID 1885595.
- 1 2 "NCBI (National Center for Biotechnology Information)".
- ↑ "National Center for Biotechnology Information".
- 1 2 "NCBI: Nucleotide". National Center for Biotechnology Information. Retrieved 2012-04-29.
- ↑ "Softberry: FGENES". Softberry, Inc. Retrieved March 31, 2012.
- 1 2 "NCBI:MapViewer". NCBI. Retrieved 2012-03-30.
- ↑ "EST Profile Viewer".
- ↑ "GeneNote".
- 1 2 Brendel V, Bucher P, Nourbakhsh IR, Blaisdell BE, Karlin S (March 1992). "Methods and algorithms for statistical analysis of protein sequences". Proc. Natl. Acad. Sci. U.S.A. 89 (6): 2002–6. doi:10.1073/pnas.89.6.2002. PMC 48584
 . PMID 1549558.
. PMID 1549558. - 1 2 "Biology WorkBench 3.2". San Diego Supercomputer Center. Retrieved April 20, 2012.
- ↑ Petersen TN, Brunak S, von Heijne G, Nielsen H (2011). "SignalP 4.0: discriminating signal peptides from transmembrane regions". Nat. Methods. 8 (10): 785–6. doi:10.1038/nmeth.1701. PMID 21959131.
- ↑ Oppermann U, Kavanagh K, Guo K, Ng SS, Lukacik P, Wu X, Dubinina E, Shafqat N, Bray J, Marsden BD, Sharma S, Vedadi M, Delft FV, Sundstrom M (2005). "SDR goes SGC: a Structural Genomics Initiative". In H Weiner, B Plapp, R Lindahl, E Maser. Enzymology and Molecular Biology of Carbonyl Metabolism. 12. West Lafayette, Ind: Purdue University Press. pp. 235–241. ISBN 1-55753-384-9.
- ↑ Kerrien S, Aranda B, Breuza L, Bridge A, Broackes-Carter F, Chen C, Duesbury M, Dumousseau M, Feuermann M, Hinz U, Jandrasits C, Jimenez RC, Khadake J, Mahadevan U, Masson P, Pedruzzi I, Pfeiffenberger E, Porras P, Raghunath A, Roechert B, Orchard S, Hermjakob H (January 2012). "The IntAct molecular interaction database in 2012". Nucleic Acids Res. 40 (Database issue): D841–6. doi:10.1093/nar/gkr1088. PMC 3245075
 . PMID 22121220.
. PMID 22121220. - ↑ Stelzl U, Worm U, Lalowski M, Haenig C, Brembeck FH, Goehler H, Stroedicke M, Zenkner M, Schoenherr A, Koeppen S, Timm J, Mintzlaff S, Abraham C, Bock N, Kietzmann S, Goedde A, Toksöz E, Droege A, Krobitsch S, Korn B, Birchmeier W, Lehrach H, Wanker EE (September 2005). "A human protein-protein interaction network: a resource for annotating the proteome". Cell. 122 (6): 957–68. doi:10.1016/j.cell.2005.08.029. PMID 16169070.
- ↑ Sowa ME, Bennett EJ, Gygi SP, Harper JW (July 2009). "Defining the human deubiquitinating enzyme interaction landscape". Cell. 138 (2): 389–403. doi:10.1016/j.cell.2009.04.042. PMC 2716422
 . PMID 19615732.
. PMID 19615732. - 1 2 "Basic Local Alignment Search Tool". National Center for Biotechnology Information. Retrieved March 3, 2012.
- ↑ Thompson JD, Higgins DG, Gibson TJ (November 1994). "CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice". Nucleic Acids Res. 22 (22): 4673–80. doi:10.1093/nar/22.22.4673. PMC 308517
 . PMID 7984417.
. PMID 7984417. - ↑ Honeth G, Bendahl PO, Ringnér M, Saal LH, Gruvberger-Saal SK, Lövgren K, Grabau D, Fernö M, Borg A, Hegardt C (2008). "The CD44+/CD24- phenotype is enriched in basal-like breast tumors". Breast Cancer Res. 10 (3): R53. doi:10.1186/bcr2108. PMC 2481503
 . PMID 18559090.
. PMID 18559090. - ↑ Kuby J, Kindt TJ, Goldsby RA, Osborne BA (2007). Kuby immunology (6th ed.). San Francisco: W.H. Freeman. pp. Appendix A, 3–4. ISBN 1-4292-0211-4.